Access GW data with ODA_API#
[ ]:
# oda_api should be installed with GW support
%pip install oda_api[gw]
[1]:
from oda_api.api import DispatcherAPI
from matplotlib import pyplot as plt
plt.rcParams['figure.figsize'] = (10, 7)
[2]:
dispatcher_url = 'https://www.astro.unige.ch/mmoda/dispatch-data'
disp=DispatcherAPI(url=dispatcher_url, instrument='mock')
Query events in the time range (catalog and skymap)#
[3]:
par_dict={
"T1": "2019-07-27T06:03:29.000",
"T2": "2019-08-27T06:03:44.000",
"contour_levels": "50,90",
"do_cone_search": "false",
"instrument": "gw",
"product": "gw_skymap_image",
"product_type": "Real",
}
data_collection = disp.get_product(**par_dict)
please beware that by default, in a typical setup, oda_api will not output much. To learn how to increase the verbosity, please refer to the documentation: https://oda-api.readthedocs.io/en/latest/user_guide/ScienceWindowList.html?highlight=logging#Let's-get-some-logging .
To disable this message you can pass `.get_product(..., silent=True)`
[4]:
data_collection.show()
ID=0 prod_name=dispatcher_catalog_0 meta_data:
ID=1 prod_name=skymap_GW190727_060333_1 meta_data: {}
ID=2 prod_name=skymap_GW190728_064510_2 meta_data: {}
ID=3 prod_name=skymap_GW190731_140936_3 meta_data: {}
ID=4 prod_name=skymap_GW190803_022701_4 meta_data: {}
ID=5 prod_name=skymap_GW190814_5 meta_data: {}
ID=6 prod_name=gw_skymap_image_6 meta_data:
Event catalog#
[5]:
data_collection.dispatcher_catalog_0.table
[5]:
Table length=5
| index | commonName | GPS | luminosity_distance | luminosity_distance_lower | luminosity_distance_upper | mass_1_source | mass_1_source_lower | mass_1_source_upper | mass_2_source | mass_2_source_lower | mass_2_source_upper |
|---|---|---|---|---|---|---|---|---|---|---|---|
| int64 | str15 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 |
| 0 | GW190727_060333 | 1248242632.0 | 3300.0 | -1500.0 | 1540.0 | 38.0 | -6.2 | 9.5 | 29.4 | -8.4 | 7.1 |
| 1 | GW190728_064510 | 1248331528.5 | 870.0 | -370.0 | 260.0 | 12.3 | -2.2 | 7.2 | 8.1 | -2.6 | 1.7 |
| 2 | GW190731_140936 | 1248617394.6 | 3300.0 | -1720.0 | 2390.0 | 41.5 | -9.0 | 12.2 | 28.8 | -9.5 | 9.7 |
| 3 | GW190803_022701 | 1248834439.9 | 3270.0 | -1580.0 | 1950.0 | 37.3 | -7.0 | 10.6 | 27.3 | -8.2 | 7.8 |
| 4 | GW190814 | 1249852257.0 | 240.0 | -50.0 | 40.0 | 23.2 | -1.0 | 1.1 | 2.59 | -0.09 | 0.08 |
[6]:
data_collection.dispatcher_catalog_0.table.meta
[6]:
{'data': 'GW in time interval',
't1': 1248242627,
't2': 1250921042,
'FRAME': None,
'COORD_UNIT': None,
'LON_NAME': None,
'LAT_NAME': None}
Skymaps#
[7]:
skmap = data_collection.skymap_GW190727_060333_1
skmap.show()
------------------------------
name: skymap_GW190727_060333
meta_data dict_keys([])
number of data units 1
------------------------------
data uniti 0 ,name:
Skymaps are encoded in Healpix MOC format
[8]:
skmap_data_unit = skmap.get_data_unit(0)
skmap_data_unit.header
[8]:
{'BITPIX': 8,
'COORDSYS': 'C',
'CREATOR': 'ligo-skymap-from-samples',
'DATE': '2021-01-26T20:10:21.896462',
'DATE-OBS': '2019-07-27T06:03:34.000000',
'DISTMEAN': 3300.149492134,
'DISTSTD': 916.3541042391864,
'GCOUNT': 1,
'HISTORY': 'or_samples /local/condor/execute/dir_24620/tmp74es3tsj.h5',
'INDXSCHM': 'EXPLICIT',
'MJD-OBS': 58691.25247685185,
'MOCORDER': 9,
'NAXIS': 2,
'NAXIS1': 40,
'NAXIS2': 16896,
'ORDERING': 'NUNIQ',
'ORIGIN': 'LIGO/Virgo',
'PCOUNT': 0,
'PIXTYPE': 'HEALPIX',
'TFIELDS': 5,
'TFORM1': 'K',
'TFORM2': 'D',
'TFORM3': 'D',
'TFORM4': 'D',
'TFORM5': 'D',
'TTYPE1': 'UNIQ',
'TTYPE2': 'PROBDENSITY',
'TTYPE3': 'DISTMU',
'TTYPE4': 'DISTSIGMA',
'TTYPE5': 'DISTNORM',
'TUNIT2': 'sr-1',
'TUNIT3': 'Mpc',
'TUNIT4': 'Mpc',
'TUNIT5': 'Mpc-2',
'VCSVERS': 'ligo.skymap 0.5.0',
'XTENSION': 'BINTABLE'}
and can be saved as fits
[9]:
skmap.write_fits_file('skymap_GW190727_060333.fits')
Contours#
[10]:
cont = data_collection.gw_skymap_image_6
Plot contours for all events
[11]:
from oda_api.plot_tools import OdaGWContours
[12]:
oda_cont = OdaGWContours(cont)
[13]:
_ = oda_cont.show()
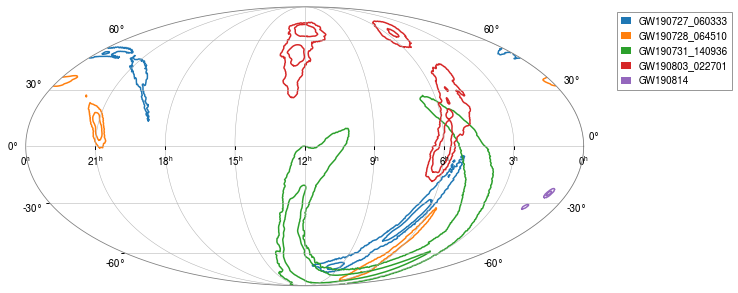
or for single event
[14]:
_ = oda_cont.show('GW190731_140936')
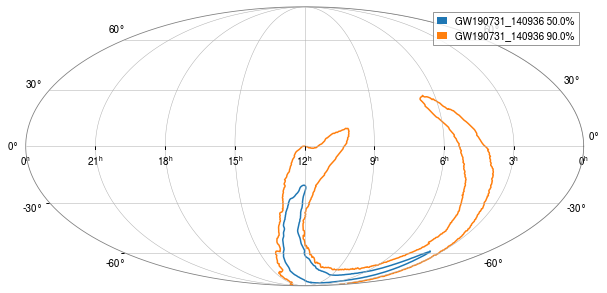
Spectrogram for the time range#
[15]:
par_dict={
"T1": "2017-08-17T12:40:54",
"T2": "2017-08-17T12:41:10",
"T_format": "isot",
"detector": "H1",
"instrument": "gw",
"product": "gw_spectrogram",
"qmax": 64,
"qmin": 4,
"product_type": "Real",
"whiten": True
}
data_collection = disp.get_product(**par_dict)
please beware that by default, in a typical setup, oda_api will not output much. To learn how to increase the verbosity, please refer to the documentation: https://oda-api.readthedocs.io/en/latest/user_guide/ScienceWindowList.html?highlight=logging#Let's-get-some-logging .
To disable this message you can pass `.get_product(..., silent=True)`
/home/dsavchenko/Projects/cloud/dispatcher-container/oda_api/oda_api/api.py:932: UserWarning:
----------------------------------------------------------------------------
the parameter: T_format is not among valid ones:
['src_name', 'RA', 'DEC', 'T1', 'T2', 'token', 'detector', 'whiten', 'qmin', 'qmax']
this will throw an error in a future version
and might break the current request!
----------------------------------------------------------------------------
warnings.warn(msg)
[16]:
data_collection.show()
ID=0 prod_name=Spectrogram_0 meta_data:
[17]:
sgram = data_collection.Spectrogram_0
produces Spectrogram object of gwpy library
[18]:
print(sgram)
Spectrogram([[1.45331742e-02, 1.45331742e-02, 1.81692112e-02,
..., 2.41175323e-04, 2.73469894e-04,
2.73469894e-04],
[1.63602121e-02, 1.63602121e-02, 2.03857105e-02,
..., 8.98046201e-05, 5.88277908e-05,
5.88277908e-05],
[1.72100142e-02, 1.72100142e-02, 2.16290895e-02,
..., 3.00785730e-04, 2.87213072e-04,
2.87213072e-04],
...,
[1.61051437e-01, 1.61051437e-01, 1.73868716e-01,
..., 5.05741846e-05, 1.13808557e-04,
1.13808557e-04],
[2.14594558e-01, 2.14594558e-01, 2.33802274e-01,
..., 1.66363490e-04, 2.15521693e-04,
2.15521693e-04],
[2.79544115e-01, 2.79544115e-01, 3.06449860e-01,
..., 2.13164400e-04, 2.63086869e-04,
2.63086869e-04]]
unit: s,
name: Spectrogram,
epoch: 1187008872.0,
channel: None,
x0: 1187008872.0 s,
dx: 0.016 s,
xindex: [1.18700887e+09 1.18700887e+09 1.18700887e+09 ... 1.18700889e+09
1.18700889e+09 1.18700889e+09] s,
y0: 22.50790790392765 Hz,
dy: None,
yindex: [ 22.5079079 22.69130096 22.87618828 ... 1270.26857957
1280.61865 1291.05305217] Hz)
[19]:
_ = sgram.plot()
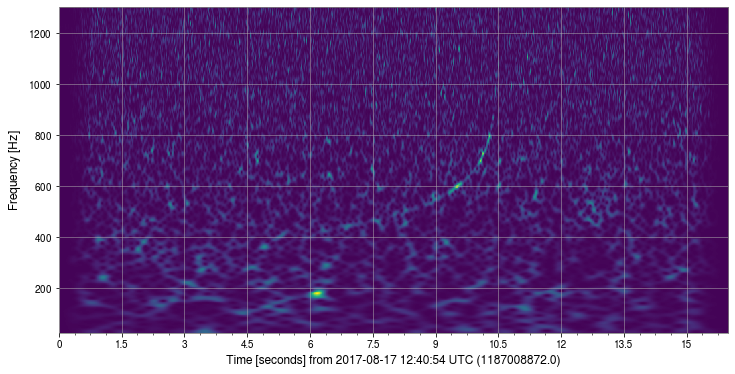
Strain time series#
[20]:
par_dict={
"T1": "2017-08-17T12:41:02",
"T2": "2017-08-17T12:41:06",
"T_format": "isot",
"detector": "H1",
"instrument": "gw",
"product": "gw_strain",
"fmax": 400,
"fmin": 30,
"product_type": "Real",
"whiten": True
}
data_collection = disp.get_product(**par_dict)
please beware that by default, in a typical setup, oda_api will not output much. To learn how to increase the verbosity, please refer to the documentation: https://oda-api.readthedocs.io/en/latest/user_guide/ScienceWindowList.html?highlight=logging#Let's-get-some-logging .
To disable this message you can pass `.get_product(..., silent=True)`
/home/dsavchenko/Projects/cloud/dispatcher-container/oda_api/oda_api/api.py:932: UserWarning:
----------------------------------------------------------------------------
the parameter: T_format is not among valid ones:
['src_name', 'RA', 'DEC', 'T1', 'T2', 'token', 'detector', 'whiten', 'fmin', 'fmax']
this will throw an error in a future version
and might break the current request!
----------------------------------------------------------------------------
warnings.warn(msg)
Produces both original and whitened-bandpassed strain time series
[21]:
data_collection.show()
ID=0 prod_name=Strain_0 meta_data:
ID=1 prod_name=Strain_bp_1 meta_data:
[22]:
bp_strain = data_collection.Strain_bp_1
[23]:
print(bp_strain)
TimeSeries([-0.01020188, 0.02356718, 0.05186042, ...,
0.03916119, 0.02171726, -0.00103156]
unit: dimensionless,
t0: 1187008880.0 s,
dt: 0.000244140625 s,
name: Strain_bp,
channel: None)
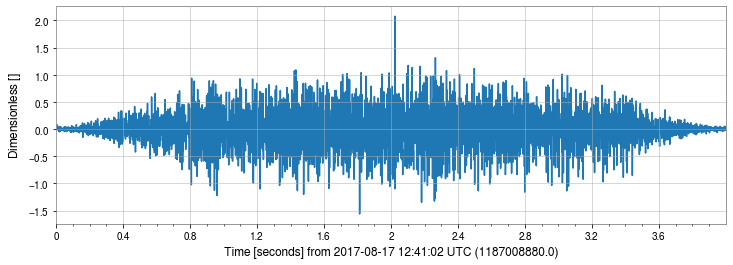
[24]:
_ = bp_strain.plot()